Biodegradation Potential of Indigenous Soil Bacteria Against Chlorpyrifos in Suthamalli Reservoir, Tirunelveli, India.
1
Department of Environmental Studies,
School of Energy, Environment and Natural Resources,
Madurai Kamaraj University,
Madurai,
Tamil Nadu
India
Corresponding author Email: roshanfathima1998@gmail.com
DOI: http://dx.doi.org/10.12944/CWE.20.3.21
Copy the following to cite this article:
Fathima M. M. M, Kannan S, Samy D. P. Biodegradation Potential of Indigenous Soil Bacteria Against Chlorpyrifos in Suthamalli Reservoir, Tirunelveli, India. Curr World Environ 2025;20(3). DOI:http://dx.doi.org/10.12944/CWE.20.3.21
Copy the following to cite this URL:
Fathima M. M. M, Kannan S, Samy D. P. Biodegradation Potential of Indigenous Soil Bacteria Against Chlorpyrifos in Suthamalli Reservoir, Tirunelveli, India. Curr World Environ 2025;20(3).
Download article (pdf)
Citation Manager
Publish History
Select type of program for download
| Endnote EndNote format (Mac & Win) | |
| Reference Manager Ris format (Win only) | |
| Procite Ris format (Win only) | |
| Medlars Format | |
| RefWorks Format RefWorks format (Mac & Win) | |
| BibTex Format BibTex format (Mac & Win) |
Article Publishing History
| Received: | 2025-07-31 |
|---|---|
| Accepted: | 2025-09-11 |
| Reviewed by: | 
 Eman Hashim
Eman Hashim
|
| Second Review by: |

 Jayath Kirthisinghe
Jayath Kirthisinghe
|
| Final Approval by: | Dr. R K Aggarwal |
Introduction
Chlorpyrifos (CPF) is a widely used organophosphate pesticide that is commonly applied in rural areas to control pests in crops such as paddy, banana, and vegetables. However, it persists in soil and water, breaking down into harmful by-products such as 2-hydroxy-3,5,6-trichloropyridine, this leads to serious environmental and health issues. Over time, CPF can accumulate in rural environments, affecting soil and water quality as well as non-target organisms. Traditional methods for cleaning up CPF, such as chemical and physical treatments, are often expensive and ineffective. As a result, microbial degradation has emerged as a promising, eco-friendly, and cost-efficient alternative.2
Many studies have focused on isolating microbes that can break down CPF from contaminated sites. Bacterial genera such as Bacillus, Pseudomonas sp., Klebsiella sp., and Acinetobacter are particularly effective, as they utilize enzymes like organophosphate hydrolases and phosphodiesterase to degrade CPF, potentially leading to complete mineralization.1,5 Additionally, bacterial consortia have been found to work together in a manner that accelerates CPF degradation, following first-order kinetics.4
In the Indian context, indigenous bacterial isolates and optimized microbial consortia have shown high resistance and efficiency in degrading CPF, suggesting that using locally adapted strains may be beneficial for bioremediation efforts.6
These site-specific microbial resources are essential in flooded agricultural areas where pesticide use is high. The Suthamalli Store in Tirunelveli district, Tamil Nadu, serves as a key water source for flood-affected crops such as paddy, black gram, and banana.
The heavy use of pesticides in surrounding areas increases the likelihood of CPF accumulating in soil and sediment. Identifying and characterizing local CPF-degrading bacteria from this source soil could offer an effective natural solution for reducing pesticide pollution. Therefore, this study aims to (i) isolate CPF-degrading bacterial strains from the soil of the Suthamalli Store, (ii) characterize their taxonomic profile and degradation potential, and (iii) evaluate their suitability for sustainable bioremediation approaches.
Materials and Methods
Study area
Suthamalli Reservoir is located at 11°03'53'' N latitude and 79°13'35'' E longitude. Suthamalli Reservoir was built primarily to provide water for domestic and agricultural uses. Banana, black gram, and paddy fields are the most cultivable vegetation around the reservoir. The agricultural land near the reservoir measures 2556.69 hectares. The Suthamalli reservoir is a subbasin of the Thamirabarani River, which flows from the Agasthiyar peak of the Pothigai hills in the Western Ghats and empties into the Gulf of Mannar.
Pesticide and Chemicals
Commercial-grade chlorpyrifos (Dursban, EC 20%), an organophosphate pesticide, was acquired for this study. It was purchased at a neighboring agrochemical store in the Tirunelveli area of Tamil Nadu. The analytical grade chemicals were purchased from HIMEDIA in Chennai.
Experimental Design
Sample Collection
In the Tamil Nadu district of Tirunelveli, soil samples were taken from the Suthamalli Reservoir's. The Chlorpyrifos pesticide is frequently used in the banana field to remove the pests and thus the pesticide residues were flow into the reservoir through the runoff. After removing the soil's surface of any residual debris from plants, grasses, and trash, the samples had been collected. Shovel was used to dig the soil and 10-15 cm deep soil was collected as a sample in clean and sterilized plastic bottles. Two distinct locations were used to collect the sample i.e. site 1 and site 2 from the banana field in sterilized bottles which were labeled with date, time and sample location as SR-1, SR -2. Samples were kept in a cold box containing ice and transported to the Environmental studies laboratory of Madurai Kamaraj University. Then the sample was air dried in the desk for 24 hours and stock in a icebox at 4oC for microbial study.
Sample preparation
The soil samples were taken from the fridge and big lumps were broken into fine powder. Soil sample was sieved through 200 micrometer sieve and the collected fine particles of soil as sample for isolation of bacteria tolerant to Chlorpyrifos pesticide.
Pesticide residue analysis employed the solid–liquid extraction combined with low-temperature purification method (SLE-LTP)
SLE-LTP involves separation of chemicals into organic phase- acetonitrile and other contaminants soluble in aqueous phase like dust, organic matter (eg. Plant residues) in aqueous phase. Aqueous phase consists of water which solidifies at -20°C and traps the water-soluble compounds and acetonitrile dissolves the chemical compounds like pesticides, which can be separated after 1 hr. The extracted acetonitrile is treated with anhydrous sodium sulfate to eliminate any remaining water. Samples can then be evaluated using a variety of qualitative and quantitative methods.7
Extraction of Pesticides from Soil
The 50 ml centrifuge tube was filled with four ml of soil sample and four ml of distilled water, followed by eight ml of acetonitrile. The material was homogenized using a vortex for 5 minutes before being stored at -20oC for 1.5 hours. Two layers were observed, one was an aqueous layer at the bottom having soil at bottom and the other was an organic layer at the top. Organic layer consisting of acetonitrile was transferred to another 50 ml centrifugation tube and 0.375g of anhydrous sodium sulphate was added to it and the sample was subjected to a vortex for 30 seconds to remove the water from the sample. The sample was then centrifuged at 4000 rpm for ten minutes. The resulting solution was transferred to another 50 mL centrifuge tube and dried in beakers covered with holes in aluminum foil for 4-6 days. Dried samples of beakers are used for UV-Visible spectroscopy, TLC, FTIR and GC-MS analysis.8
Analytical Methods
UV- Visible Spectroscopy Analysis
Three ml of the extracted pesticide from soil sample by SLE- LTP method, was analyzed in the UV- Visible spectroscopy in 600 nm. Acetonitrile HPLC grade used as the control.
Gas Chromatography - Mass Spectrometry Analysis
A Shimadzu 2030 GC equipped with mass spectrometry was used to analyze the pesticides and its residues in the soil sample. The Sample was extracted in Acetonitrile HPLC grade solvent. The oven temperature was kept at one hundred degrees Celsius for five minutes before being increased to 300°C for 20 minutes. The detector temperature was 300 degrees Celsius. Nitrogen was used as a carrier gas. The chemicals were identified using mass spectra, and the results were compared using the National Institute of Standards and Technology (NIST) collection.8
Isolation of Bacteria
Soil bacteria were isolated via the standard and serial dilution method, with one gram of soil from each site suspended in 9 ml of sterile (double distillation water). The sequential dilution method was utilized up to a 10-7 dilution. A small aliquot (0.1ml) of each dilution (10-1, 10-2, 10-3, 10-4, 10-5, 10-6, and 10-7) was obtained and streaked over nutrient agar plates to obtain bacterial colonies. After 24 hours of incubation at 34oC, isolated colonies were selected and pure cultures grew on nutrient agar plates.9
Identification of Bacteria
To characterize the physical characteristics of the bacterial isolates from the soil, microscopic and biochemical investigations were carried out. They were checked for pigmentation, colony morphology, and standard preliminary tests including Gram's staining and biochemical assays that were validated by Bergey's manual of determinative bacteriology.
Biodegradation study
In a sterilized laminar air flow chamber, one milliliter of bacterial broth culture was inoculated into 100 milliliters of nutrient broth and 5 milliliters of 50 uL/ml of chlorpyrifos pesticide. The culture was kept in a growth chamber for ten days at 37°C. The degradation was determined by monitoring the absorbance of the 3 ml aliquot of sample at 320 nm at an interval of 24 hrs in UV - visible spectroscopy.10,11
Growth Efficacy Study
The growth performance of bacterial isolates in the presence of chlorpyrifos was assessed in liquid Minimal Salt Medium (MSM) supplemented with chlorpyrifos (50 µg/mL). Each 100 mL culture was infused with 1 mL of bacterial broth and cultured at 37 °C in a shaking incubator (150 rpm) for 10 days. Control flasks included: (C1) uninoculated MSM + chlorpyrifos (to monitor abiotic loss) and (C2) MSM + bacteria without pesticide (to monitor background growth). Growth was monitored at 24 h intervals by measuring optical density at 600 nm (OD600), and CFU counts were performed at selected intervals to validate bacterial growth.12
Results
Chlorpyrifos has been an efficient pesticide in agriculture, its detrimental impacts on the health of people and the environment have prompted more regulations and a move toward safer pest management techniques. Microorganisms found in the soil of chlorpyrifos-infected banana fields may break down this pesticide. Numerous harmful environmental consequences result from the excessive application of these organophosphate pesticides on banana fields. It is important to eliminate contaminants from the environment based on their level of toxicity. The most common way of removing pesticides is biologically, as soil microorganisms may metabolize these toxic substances into non- toxic substances.
Extract the pesticide from the soil and water sample by SLE-LTP method of suthamalli reservoir, in which at 288 nm the peak of chlorpyrifos is observed in the soil sample by UV-Visible spectrophotometer.
The UV-visible absorption spectrum of Chlorpyrifos pesticide demonstrates a characteristic strong absorbance peak at 225 nm, distinguishing it clearly from the soil sample spectrum. The soil sample shows moderate absorbance in the same region, which may indicate the presence of trace organic compounds or pesticide residues. The comparative profile confirms that Chlorpyrifos can be reliably detected and quantified in the UV range of 220–285 nm. Beyond 300 nm, spectral overlap diminishes due to negligible absorption. This peak indicates preliminary the presence of Chlorpyrifos residues in collected soil sample.
The extracted chlorpyrifos pesticide from the soil sample was identified by GCMS using the NIST library. Figure 1 showed that chromatogram image reveals a clear separation, three distinct analytes eluting between 5 and 17 minutes, and a solvent front at the start. The data can be further analyzed to identify substances based on mass spectral information.
Upon GCMS analysis shows in a (Table 1) a retention time of 12.434 min in the 3rd peak and 22.149 min in 5th peak as chlorpyrifos and Phosphoric acid, tris (2,4-di-tert-butylphenyl) ester respectively, which was used as a stabilizer and an antioxidant. The obtained soil sample encompassed chlorpyrifos, as verified by these findings.
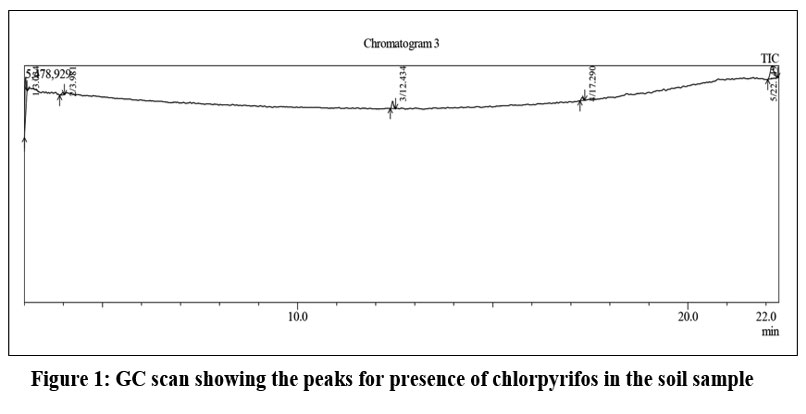 | Figure 1: GC scan showing the peaks for presence of chlorpyrifos in the soil sample
|
Table 1: GC–MS Characterization of Environmental Contaminants and Additives
Peak ID | Retension Time (Min) | Initial Time (Min) | Final Time (Min) | Peak Area (counts.s-1) | Relative Peak Area (%) | Peak Height (Counts) | Base lon (m/z) | Identified Compound |
1 | 3.034 | 3.005 | 3.070 | 2219622 | 44.54 | 914872 | 28.05 | Nickel tetracarbonyl |
2 | 3.981 | 3.905 | 4.025 | 184733 | 3.71 | 53723 | 1.95 | Hydrogen |
3 | 12.434 | 12.375 | 12.510 | 513835 | 10.31 | 170278 | 96.95 | Chlorpyrifos |
4 | 17.290 | 17.235 | 17.355 | 239177 | 4.80 | 73127 | 149.00 | Bis(2-ethylhexyl) phthalate |
5 | 22.149 | 22.045 | 22.300 | 1826415 | 36.65 | 297797 | 647.45 | Tris(2,4-di-tert-butylphenyl) phosphate |
4983782 | 100.00 | 1509797 |
Chlorpyrifos (CAS No. 2921-88-2) is an organophosphate insecticide with the molecular formula C9H11Cl3NO3PS and a molecular weight of 349 g·mol-1 . It is also commercially known under trade names such as Dursban. The compound is systematically named as O,O-diethyl O-(3,5,6-trichloro-2-pyridyl) phosphorothioate, according to IUPAC nomenclature.
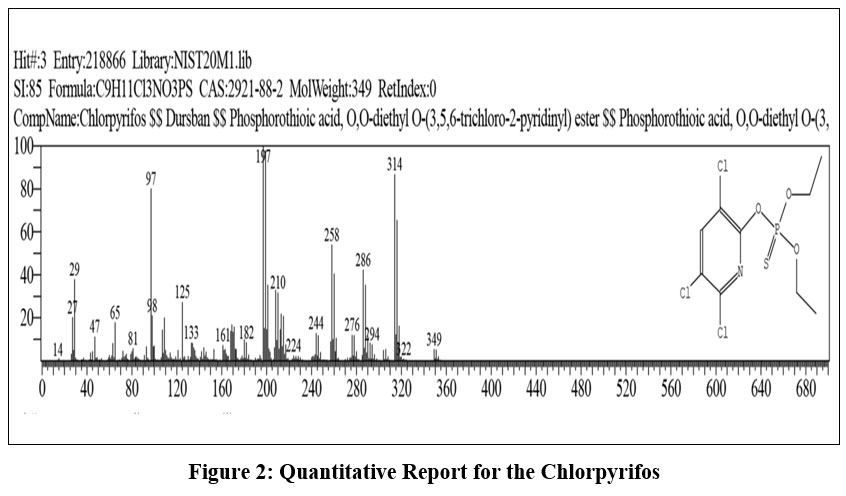 | Figure 2: Quantitative Report for the Chlorpyrifos
|
Tris (2,4-di-tert-butylphenyl) phosphate (CAS No. 95906-11-9) is an organophosphate compound with the molecular formula C42H63O4P and a molecular weight of 662 g·mol-1 . This compound is also referred to as Phenol, 2,4-bis(1,1-dimethylethyl)-, phosphate or Phenol, 2,4-bis(1,1-dimethylethyl)-, 1,1',1''-phosphate, depending on the nomenclature system used. It is widely recognized as an antioxidant additive in polymers due to its thermal stability and resistance to oxidative degradation.
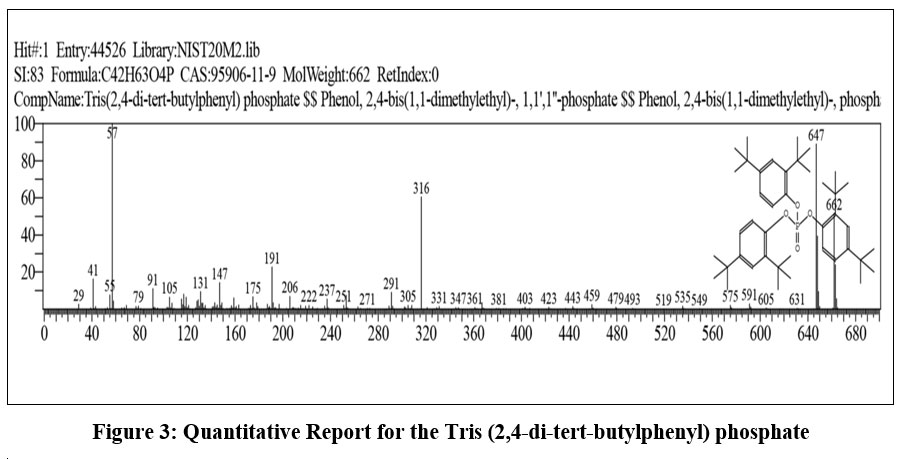 | Figure 3: Quantitative Report for the Tris (2,4-di-tert-butylphenyl) phosphate
|
Isolation of chlorpyrifos pesticide degrading bacterial isolates
The soil sample from a banana field containing chlorpyrifos pesticide included all eight of the bacterial isolates that were able to use organophosphate pesticide as their only source of carbon for growth and development. They were presumptiviely identified according to their physical and biochemical characteristics as presented in the (Table 2.). The RFS1, RFS2, RFS3, RFS4, RFS5, RFS6, RFS7 and RFS8 isolates were identified as Streptococcus sp, Arthrobacter sp, Enterobacter sp, Rhodococcus sp, Staphylococcus sp, Bacillus sp, Pseudomonas sp and Proteus sp respectively.
Table 2: Phenotypic and Biochemical Characteristics of Bacterial Isolates
S.no | Shape | Gram Staining | Indole | Methyl red | Voges Proskauer | Citrate | Name of the species |
RFS1 | Spherical shape | Gram Positive | -ve | -ve | -ve | -ve | Streptococcus sp |
RFS2 | Rod shape | Gram Positive | -ve | -ve | -ve | -ve | Arthobacter sp |
RFS3 | Rod shape | Gram Negative | -ve | -ve | +ve | +ve | Enterobacter sp |
RFS4 | Cocci shape | Gram Positive | -ve | -ve | -ve | -ve | Rodococcus sp |
RFS5 | Spherical & grape-like clusters | Gram Positive | -ve | -ve | -ve | +ve | Staphylococcus sp |
RFS6 | Rod shape | Gram Positive | -ve | -ve | +ve | +ve | Bacillus sp |
RFS7 | Rod shape | Gram Positive | - ve | -ve | -ve | +ve | Pseudomonas sp |
RFS8 | Rod shape | Gram Negative | -ve | +ve | -ve | +ve | Proteus sp. |
Bacterial isolates RFS1, RFS4 and RFS6 were observed that they are be tolerant in pesticide tolerance test so further they were used for bio-degradation study for monitoring their ability to degrade chlorpyrifos. significant changes in the growth curve (observed at 600 nm) of the bacteria were observed in addition to chlorpyrifos to the bacterial culture. The addition of chlorpyrifos insecticide to the culture medium increases bacterial growth because it gives an additional carbon source.
Each isolate's optical density at 600 nm was measured at various time intervals. i.e., 24, 48, 72, 96, 120, 144, 168, 192, 216 and 264 on nutrient broth. Out of 8 isolated bacterial strains only 3 isolates-RFS1, RFS4 and RFS6 were selected on their ability to grow in different concentration of chlorpyrifos (25,50,75 and100uL/ml) for the bio-degradation study shown in (Table 3). The outcome of the isolates' growth reaction while chlorpyrifos existed showed in the table 3 RFS1, RFS4 and RFS6 named as Streptococcus, Rodococcus and Bacillus were utilized the pesticide as the only carbon and energy source. Streptococcus showed maximum growth (OD=1.936) (Fig 4.) and Bacillus (OD= 1.842) (Fig 5.) in 10th days. The same results was found that Bacillus tropicus degrade 60% chlorpyrifos after 48 hrs. of incubation found by Rayu et al.5 while Rodococcus recorded highest growth in 9th days incubation (Fig 6). that Chlorpyrifos was subjected to biodegradation study with RFS1, RFS4 and RFS6 bacterial isolates for a period of 12 days. In the light of this fact, biodegradation especially microbial degradation has proven to be a suitable method for pesticide elimination. Most of the previous study indicated that Rhodobacter could effectively degrade the pesticide like 2,4- D and chlorpyrifos.3 Chlorpyrifos is changed into chlorpyrifos oxon by the oxidation reaction, which is one of the steps involved in the mechanism of chlorpyrifos degradation.
Table 3: Growth curve and Biodegradation study for chlorpyrifos using RFS1, RFS4 and RFS6 bacterial culture
Time | RFS1 and chlorpyrifos | RFS1 control | Chlorpyrifos with RFS1 | RFS4 and chlorpyrifos | RFS4 control | Chlorpyrifos with RFS4 | RFS6 and chlorpyrifos | RFS6 control | Chlorpyrifos with RFS6 |
0 | 0 | 0 | 1.611 | 0 | 0 | 1.441 | 0 | 0 | 1.441 |
24 | 0.648 | 0.108 | 1.507 | 0.522 | 0.213 | 1.325 | 0.584 | 0.192 | 1.405 |
48 | 0.748 | 0.153 | 1.732 | 0.636 | 0.245 | 1.391 | 0.668 | 0.201 | 1.628 |
72 | 0.721 | 0.241 | 1.596 | 0.609 | 0.267 | 1.486 | 0.697 | 0.245 | 1.523 |
96 | 0.776 | 0.366 | 1.724 | 0.623 | 0.301 | 1.491 | 0.701 | 0.312 | 1.673 |
120 | 0.799 | 0.388 | 1.776 | 0.627 | 0.367 | 1.507 | 0.733 | 0.345 | 1.687 |
144 | 0.829 | 0.393 | 1.81 | 0.63 | 0.345 | 1.539 | 0.714 | 0.367 | 1.685 |
168 | 0.838 | 0.418 | 1.873 | 0.619 | 0.387 | 1.652 | 0.696 | 0.421 | 1.711 |
192 | 0.831 | 0.457 | 1.936 | 0.622 | 0.402 | 1.503 | 0.705 | 0.456 | 1.842 |
216 | 0.599 | 0.462 | 1.445 | 0.628 | 0.412 | 1.565 | 0.689 | 0.49 | 1.605 |
264 | 0.605 | 0.472 | 1.471 | 0.69 | 0.423 | 1.595 | 0.696 | 0.523 | 1.634 |
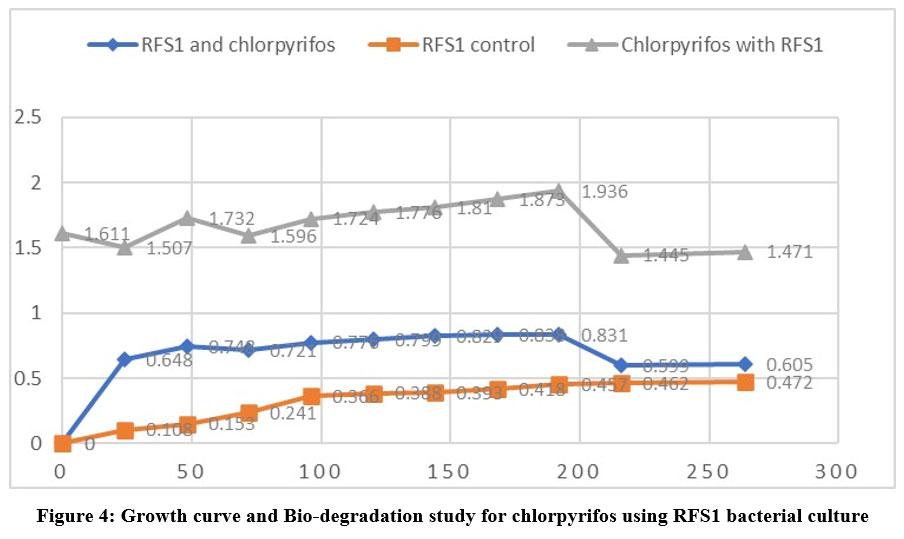 | Figure 4: Growth curve and Bio-degradation study for chlorpyrifos using RFS1 bacterial culture
|
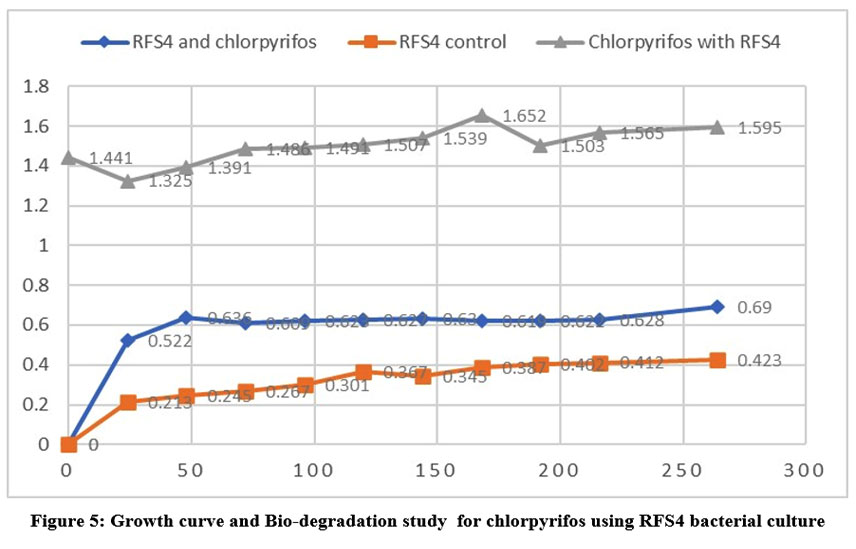 | Figure 5: Growth curve and Bio-degradation study for chlorpyrifos using RFS4 bacterial culture
|
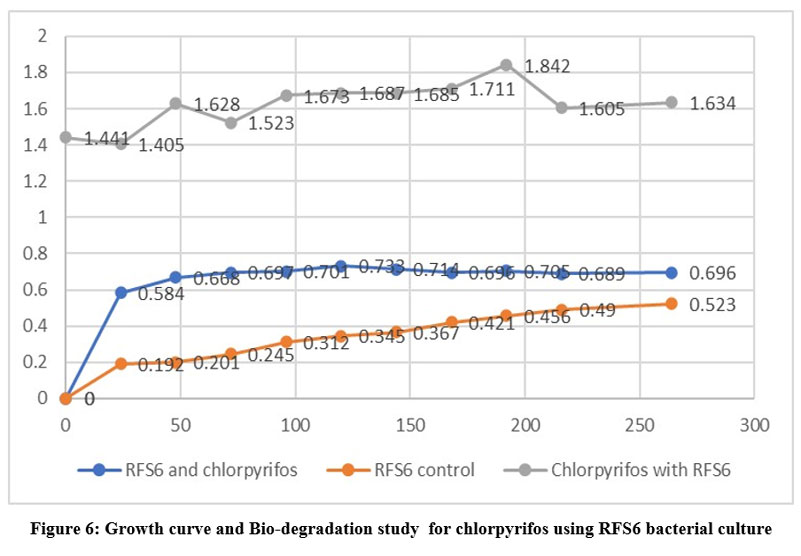 | Figure 6: Growth curve and Bio-degradation study for chlorpyrifos using RFS6 bacterial culture
|
Table 4 shows that the calculation value of growth efficacy as the relative increase in OD of the chlorpyrifos-treated culture compared to its respective control at each time point.

Table 4: Growth efficacy of bacterial cultures expressed as the relative increase in optical density (OD) of chlorpyrifos-treated samples compared to untreated controls at different time intervals.
Time (h) | RFS1 (%) | RFS4 (%) | RFS6 (%) |
0 | 0.00 | 0.00 | 0.00 |
24 | 500.00 | 145.07 | 204.17 |
48 | 388.89 | 159.59 | 232.34 |
72 | 199.17 | 128.09 | 184.49 |
96 | 112.02 | 106.98 | 124.68 |
120 | 105.93 | 70.84 | 112.46 |
144 | 110.94 | 82.61 | 94.55 |
168 | 100.48 | 59.95 | 65.32 |
192 | 81.84 | 54.73 | 54.61 |
216 | 29.65 | 52.43 | 40.61 |
264 | 28.18 | 63.12 | 33.08 |
RFS1 (Streptococcus) showed the highest initial growth efficacy (500%) at 24 h, but then gradually declined after 72 h. RFS6 (Bacillus) maintained strong growth efficacy (200–230%) up to 48 h, and still showed above 90% up to 144 h, confirming its robustness. RFS4 (Rhodococcus) had moderate efficacy (70–160%), with more stable performance across the incubation period compared to RFS1. This indicates that RFS1 is the fastest responder, but RFS6 sustains growth better over time, making it a strong candidate for long-term chlorpyrifos biodegradation.
Discussion
Chlorpyrifos, an organophosphate pesticide, has long been valued for its broad-spectrum efficacy in agriculture. However, its persistence and toxicity have raised significant concerns regarding environmental and human health, resulting in stricter regulations and growing interest in sustainable pest management strategies.18 In the current investigation, chlorpyrifos residues were found in soil samples from the Suthamalli reservoir region using UV-Visible spectroscopy and GC-MS. A distinct absorbance peak at 225 nm confirmed the pesticide’s presence in the UV spectrum, consistent with previous findings that chlorpyrifos exhibits strong UV absorption in the 220–285 nm range.16 GC–MS analysis further validated the presence of chlorpyrifos at a retention time of 12.434 min, alongside additional contaminants such as tris (2,4-Di-Tert-Butylphenyl) phosphate, indicating that pesticide residues often co-exist with industrial additives in agricultural soils.15 The isolation of eight bacterial strains capable of utilizing chlorpyrifos as the exclusive carbon source underscores the role of indigenous soil microbiome in natural detoxification processes. Among these, Streptococcus (RFS1), Rhodococcus (RFS4), and Bacillus (RFS6) exhibited significant growth responses when exposed to chlorpyrifos. Growth efficacy analyses revealed that RFS1 displayed rapid initial adaptation, reaching 500% growth efficacy at 24 h, whereas RFS6 demonstrated more sustained degradation potential, maintaining over 90% efficacy beyond 144 h. This finding is consistent with prior studies where Bacillus13 and Rhodococcus species17 were identified as efficient degraders of organophosphate due to their diverse enzymatic pathways, including hydrolysis and oxidation.14
The differential growth responses of the isolates suggest complementary degradation strategies: fast responders like Streptococcus initiate rapid breakdown, while robust degraders such as Bacillus ensure prolonged activity. These results highlight the potential of harnessing native bacterial consortia for bio-remediation of chlorpyrifos-contaminated soils. Similar microbial-mediated degradation pathways have been reported, where chlorpyrifos is oxidized to chlorpyrifos-oxon and subsequently metabolized into less toxic products.19 Thus, microbial bio-degradation represents a practical, eco-friendly strategy for pesticide removal in agricultural systems. Future work should focus on the genetic characterization of degradation enzymes and the development of bioaugmentation models for large-scale field applications.
Conclusion
Our findings’ showed the isolated bacterial strains that could use the chlorpyrifos pesticide as a soil carbon source. One important mechanism by which xenobiotic chemicals are eliminated from the environment and avoid contamination of the environment is the use of these compounds by soil bacteria. Result of this study suggest that the isolates Streptococcus, Rodococcus and Bacillus capacity to use and proliferate in the presence of chlorpyrifos. It shows that they might be applied to the bioremediation of soil and water damaged by pesticides.
Acknowledgement
Authors would like to thank Madurai Kamaraj University, Department of Environmental Studies for granting the Ph.D., research work. Author also thanks to funding agency RUSA (Rashtritya Uchchatar Shiksha Abhiyan), Madurai Kamaraj University for providing fund for a project entitled “Environmental Impact assessment of water bodies and its associated environment in selected altitudes of Southern Tamil Nadu” for two year research work 2021-2023.
Funding Sources
005/RUSA/MKU2020-2021, RUSA, MKU
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
The Manuscript incorporates all datasets produced
Ethics Statements
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Permission to reproduce material from other sources
Not Applicable
Author Contributions:
Ms. M.Mathar Fathima: Designed the experiment and methodology, writing, executed experiment, organised the data, and prepared the manuscript.
Dr.S.Kannan : Reviewed the data along with the whole manuscript.
Mrs. P.Dheivanai: Data analysis
Reference
- Akbar, S., Anwar, Z., & Irfan, M. Biodegradation of chlorpyrifos using isolates from contaminated soils. Scientific Reports.2021;11(1):22461. https://doi.org/10.1038/s41598-021-01889-9
CrossRef - Chen, S., Hu, M., & Liu, J. Insights into the microbial degradation and catalytic mechanisms of chlorpyrifos. Environmental Research.2020;184:109328. https://doi.org/10.1016/j.envres.2020.109328
CrossRef - Chen, S., Xiao, Y., & Ding, Y. A review on the microbial degradation of chlorpyrifos and its metabolite TCP. Chemosphere.2022;293:133571. https://doi.org/10.1016/j.chemosphere.2022.133571
CrossRef - Ghosh, S., Banerjee, S., & Mondal, A. Mechanism and kinetics of chlorpyrifos co-metabolism by a bacterial consortium (ERM C-1). Frontiers in Microbiology.2022;13: 881756. https://doi.org/10.3389/fmicb.2022.881756
- Rayu, S., Chandra, R., & Singh, A. Molecular characterization of chlorpyrifos-degrading bacteria from agricultural soils. Heliyon.2022;8(5):e09315. https://doi.org/10.1016/j.heliyon.2022.e09315
CrossRef - Sethunathan, N., Nair, A., & Joseph, A. Biodegradation of chlorpyrifos by an optimized Bacillus consortium isolated from Indian soils. International Journal of Pest Management.2021;67(3):205–214. https://doi.org/10.1080/09670874.2020.1777953
- Silva, F. R., Almeida, R. I., Santos, V. G., Neves, A. A., & de Queiroz, M. E. L. R. Solid–liquid extraction with low-temperature partitioning as an alternative to determine pesticide residues in teas. Journal of the Brazilian Chemical Society. 2025;36(2): 205–216. https://doi.org/10.21577/0103-5053.20240234
- Silva, T. C., Neves, A. A., Cardeal, Z. L., de Queiroz, M. E. L. R., & Heleno, F. F. Optimization and validation of the miniaturized solid–liquid extraction with low-temperature purification (SLE-LTP) method for determining fluopyram in sandy, clayey, and medium-textured soil. Journal of the Brazilian Chemical Society. 2023;34(5):673–683. https://doi.org/10.21577/0103-5053.20230013
CrossRef - Jokhakar PH, Dudhagara PR. Degradation and detoxification of monocrotophos using bacterial consortium. J App Biol Biotech.2022;10(6):127-137. DOI: 10.7324/JABB.2022.100614.
CrossRef - S. Deng. Y., Chen, D., Wang, T., et al. Rapid biodegradation of organophosphorus pesticides by Stenotrophomonas sp. G1. Journal of Hazardous Materials.2015;297:17-24. https://doi.org/10.1016/j.jhazmat.2015.04.052
CrossRef - Sharma. A. J., Pandit., Sharma R and Shirkot, P. (2016). Biodegradation résinovaran strain AST2.2 isolated from enriched cultures. Curr World Environ.2016;11:267-278. ginal Research Paper. https://doi.org/10.46488/NEPT.2020.v19i02.043
CrossRef - Kumar, G., Lal, S., Soni, S. K., Maurya, S. K., Shukla, P. K., Chaudhary, P., Bhattacherjee, A. K., & Garg, N. Mechanism and kinetics of chlorpyrifos co-metabolism by environment-restoring microbes isolated from rhizosphere of horticultural crops under subtropics. Frontiers in Microbiology. 2022;13: 891870. https://doi.org/10.3389/fmicb.2022.891870
CrossRef - Kumar Malik.. Raina Singhmar., Vivek Singh., Meenu Rathi and Vishal Ahlawat. Chlorpyrifos Degradation by Bacillus tropicus a Plant Growth Promoting Rhizobacteria. Biosciences Biotechnology Research Asia. 2024;21(1):349-354. http://dx.doi.org/10.13005/bbra/3230
CrossRef - Gupta, N., Yadav, S., & Verma, A. Microbial degradation of organophosphate pesticides: Recent advances and future prospects. Journal of Environmental Management. 2022;307:114546. https://doi.org/10.1016/j.jenvman.2022.114546
- Kumari, R., Singh, V., & Meena, R. Pesticide residues and co-contaminants in agricultural soils: Implications for soil health and microbial communities. Environmental Science and Pollution Research.2023;30(12):35029–35041.https://doi.org/10.1007/s11356-023-25688-9
- Patel, M., & Deshmukh, D. UV–Vis spectrophotometric determination of organophosphate pesticides in soil and water matrices. Environmental Technology & Innovation. 2021;24: 102000. https://doi.org/10.1016/j.eti.2021.102000
CrossRef - Rani, P., & Kumar, S. Role of Rhodococcus species in biodegradation of organophosphates: A sustainable approach for environmental detoxification. Biocatalysis and Agricultural Biotechnology. 2021;35:102082. https://doi.org/10.1016/j.bcab.2021.102082
CrossRef - Singh, A., Sharma, P., & Kaur, J. Environmental and health impacts of chlorpyrifos: Regulatory updates and microbial remediation perspectives. Ecotoxicology and Environmental Safety. 2022;239:113689. https://doi.org/10.1016/j.ecoenv.2022.113689
CrossRef - Zhang, X., Li, Y., & Chen, J. Biodegradation pathways and enzymatic mechanisms of chlorpyrifos degradation by soil microorganisms. Applied Microbiology and Biotechnology. 2020;104(21):9205–9217. https://doi.org/10.1007/s00253-020-10874-y
Abbreviations List
HPLC (High-Performance Liquid Chromatography),
GC-MS(Gas Chromatography-Mass Spectrometry),
TLC (Thin Layer Chromatography),
SLE-LTP (Solid Liquid Extraction-Low Temperature Purification),
FTIR (Fourier Transform Infrared Spectroscopy),
UV (Ultraviolet), and
NIST (National Institute of Standards and Technology).






