Bacterial Degradation of Textile Dyes by Cultures Isolated from Effluent Samples
Shilpa Pranav Tambe1
*
 , Aarfiya Shaikh1
, Aarfiya Shaikh1
 , Pranav Tambe2
, Pranav Tambe2
 , Shaikh Umair Ahmed1
, Shaikh Umair Ahmed1
 and Jaspal Kaur Oberoi1
and Jaspal Kaur Oberoi1

1
Department of Microbiology,
M.C.E Society’s Abeda Inamdar Senior College of Arts,
Science and Commerce (Autonomous), Savitribai Phule Pune University,
Pune,
Maharashtra
India
2
Department of Chemistry,
M.C.E. Society’s Abeda Inamdar Senior College of Arts, Science and Commerce (Autonomous),
Savitribai Phule Pune University,
Pune,
Maharashtra
India
Corresponding author Email: shilpatambe@azamcampus.org
DOI: http://dx.doi.org/10.12944/CWE.20.3.15
Copy the following to cite this article:
Tambe S. P, Shaikh A, Tambe P, Ahmed S. U, Oberoi J. K. Bacterial Degradation of Textile Dyes by Cultures Isolated from Effluent Samples. Curr World Environ 2025;20(3). DOI:http://dx.doi.org/10.12944/CWE.20.3.15
Copy the following to cite this URL:
Tambe S. P, Shaikh A, Tambe P, Ahmed S. U, Oberoi J. K. Bacterial Degradation of Textile Dyes by Cultures Isolated from Effluent Samples. Curr World Environ 2025;20(3).
Download article (pdf)
Citation Manager
Publish History
Select type of program for download
| Endnote EndNote format (Mac & Win) | |
| Reference Manager Ris format (Win only) | |
| Procite Ris format (Win only) | |
| Medlars Format | |
| RefWorks Format RefWorks format (Mac & Win) | |
| BibTex Format BibTex format (Mac & Win) |
Article Publishing History
| Received: | 2025-07-18 |
|---|---|
| Accepted: | 2025-12-12 |
| Reviewed by: | 
 Kiran Fatima
Kiran Fatima
|
| Second Review by: |

 Biswajit Saha
Biswajit Saha
|
| Final Approval by: | Dr. Ivana Mjhajlovic |
Introduction
Industries often release untreated wastewater into the environment, leading to pollution. Textile dyeing operations, in particular, produce vast quantities of wastewater daily on a global scale. This wastewater frequently contains hazardous chemicals, including synthetic/ noxious dyes and metal contaminants such as arsenic, cadmium, mercury, lead, and chromium, which are used in textile dye production.1 While dyes were once derived from natural sources like flowers, vegetables, roots and insects, the demand for long-lasting, intense colorants has facilitated the shift toward synthetic dyes of petrochemical origin in textile manufacturing. The preference for synthetic dyes over natural counterparts is attributed to their superior solubility in water, faster application rates, and extensive color range. In textile manufacturing, azo-based synthetic dyes are commonly chosen for their greater chemical stability than that of natural dyes.2
A dye, whether natural or synthetic, is a colored substance applied to materials such as cloth, paper, plastic, or leather to permanently alter their color. Textile industry wastewater often contains harmful synthetic colorants and toxic metallic elements, such as arsenic, cadmium, chromium, lead, and mercury. The coloring of textile fibers is achieved via chromophores present in dye molecules, while the fixation process relies on auxochromes, which form bonds with the polar sites of the fiber.3
Azo dyes represent the largest category of colorants, accounting for over 50% of all dyes used in various industries. These dyes are distinguished by their chemical structure, & this compound contains an azo linkage (–N=N–), characterized by a double bond between two nitrogen atoms, with at least one of them connected to an aromatic group. Structural attributes of azo dyes allow their classification into distinct groups: anionic dyes, non-ionic dyes, and cationic dyes. This classification comprises acid dyes, basic (cationic) dyes, direct dyes (also known as substantive dyes), disperse dyes (also known as non-ionic) dyes, as well as reactive, vat, and sulfur dyes. 4
Textile and tanning industries must carefully treat their effluents to remove any residual synthetic azo dyes, as these compounds can accumulate in the environment and pose risks of phytotoxicity and harm to aquatic ecosystems. The management of industrial wastewater is complicated by harsh physicochemical properties, such as high chemical and biological oxygen demands (COD and BOD), elevated thermal levels, high salt content and excessive amounts of suspended and dissolved solids, as well as pronounced alkalinity.5 Dye effluents can negatively impact human health through both direct and indirect exposure. They may cause a range of health issues, including nausea, hemorrhages, skin ulcers, mucous membrane irritation, kidney failure and problems with the gastrointestinal tract and liver.6 Frame shift mutations have been observed in human hepatoma cells as a result of the mutagenic effects of the dye.7 Micronuclei formation have been observed in human lymphocytes and HepG2 cells.8
Bioremediation of dye effluents is important for mitigating environmental pollution and preserving ecological balance. Various bioremediation methods have been suggested, including adsorption techniques, exposure to irradiation, coagulation and flocculation processes, chemical precipitation, membrane filtration, application of activated carbon and oxidation through chemical or electrochemical means. However, some of the above-mentioned remediation techniques require substantial Funding allocation and potentially ineffective in achieving complete dye breakdown. Dyes are xenobiotic compounds with electron-withdrawing groups, rendering them resistant to degradation. Even when degradation occurs, it often results in formation of aromatic amines that are known to be carcinogenic and mutagenic. Additionally, chemical treatments can lead to sludge formation, contributing to secondary pollution.9
Degradation of dyes through microbial or enzymatic activity offers a more environmentally friendly, cost-effective and practical solution. This approach not only breaks down the dyes but also generates less toxic by-products and produces less sludge compared to chemical treatments. Given that dye wastewater typically contains a mixture of dyes, the most effective bacterial strains are those capable of efficiently degrading multiple types of dyes.
Materials and Methods
Collection of Sample
Two locations in the Camp area (18.5258o N, 73.8806o E) of Pune region having were selected based on their significant exposure to untreated wastewater from dye manufacturing units. Liquid Samples were collected from these sites in Falcon Tubes. All the experiments were performed in triplicates and standard deviation is expressed in terms of error bar in all the graphs.
Preparation of dye stock solution
10 mg of Methyl Orange dye was weighed and dissolved in a test tube with distilled water. The volume was adjusted to 10 mL with continuous shaking, creating a stock solution for future use. A similar procedure was used for Congo Red.9
Isolation of Bacteria from Dye Effluent Sample
Microbial cultures capable of degrading dyes were screened using enrichment methods. Initially, 4 mL of collected samples were added to 10 mL of minimal medium with 0.1 mg/mL Methyl Orange dye as the sole carbon source. After incubating at 30°C for three days, 4 mL aliquots were transferred to fresh medium with the same dye concentration and incubated for another three days. This process was repeated three times over nine days for acclimatization. Following the third cycle, a 0.5 mL aliquot from samples showing complete decolorization was streaked onto Methyl Orange-enriched agar media and the plates were incubated at 30°C until visible colony formation was observed. The selected colonies were introduced into minimal medium broth with a final dye concentration of 0.1 mg/mL Methyl Orange for sub-culturing. The same procedure was applied for Congo Red.10
Decolorization Studies
A 1 mL fresh bacterial culture was inoculated into 10 mL of minimal broth with 0.1 mg/mL methyl Orange dye and maintained at 37°C in a stationary aerobic environment with maintained pH 7.0. A control without bacteria was included. After incubation, the samples were subjected to centrifugation at 4°C for 10 minutes at 10,000 rpm and decolorization was quantified by measuring absorbance at 465 nm using a UV–Vis spectrophotometer. The same procedure was applied to Congo Red dye with absorbance measured at 497 nm. Decolorization percentages were calculated using the standard formula.11

Optimization of the Decolorization process
Tests were conducted to optimize conditions for decolorizing Methyl Orange and Congo red dyes. The study examined the effects of temperature, pH, mixed dyes, single-dye consortia and mixed-dye consortia.
Effect of temperature on dye Decolorization
To assess temperature effects on decolorization, broth medium with selected strains was incubated at 30°C, 40°C and room temperature. Decolorization was measured by monitoring optical density (OD).
Effect of pH on decolorization of dye
To study different pH effects on decolorization, Methyl Orange and Congo Red dyes were incubated with bacterial cultures in broth at pH 5, 7 and 9. The pH set at the desired level by the addition of 0.1N NaOH or 0.1N HCl. Decolorization was determined using optical density (OD) at 465 nm and 497 nm.
Top of Form
Bottom of Form
Effect of Consortia on single and mix dye Decolorization
To study the effect of bacterial consortia on decolorization, Methyl Orange and Congo Red dyes were incubated with the consortia at 30°C. Decolorization was measured by optical density at 465 nm and 497 nm. For mixed dyes, the consortia were incubated with both dyes at 30°C and decolorization analysis at 465 nm using a UV–Vis spectrophotometer.
FTIR Analysis
The FTIR analysis of textile dye effluent before and after treatment was performed as it plays a vital role in characterizing molecular alterations in dye structures by monitoring changes in key functional-group vibrations after bacterial exposure. It facilitates the elucidation of biodegradation pathways through the detection of newly formed metabolites and degradation intermediates.
Results
Bacterial Strain Used
Escherichia spp., Staphylococcus spp. and Pseudomonas spp. were initially tested for their ability to biodegrade the selected dyes. These strains were sourced from the Department of Microbiology. Enterobacter spp. were isolated from textile effluent.
Biodegradation of Synthetic Dyes: Methyl Orange and Congo Red
Variations in degradation efficiency have been observed among different bacterial cultures. In our study, Pseudomonas spp. was most effective against Methyl Orange, achieving 93.42% degradation, while Escherichia spp. exhibited the highest degradation of Congo red at 88.63%. Percent degradation for other bacteria is illustrated in fig.1.
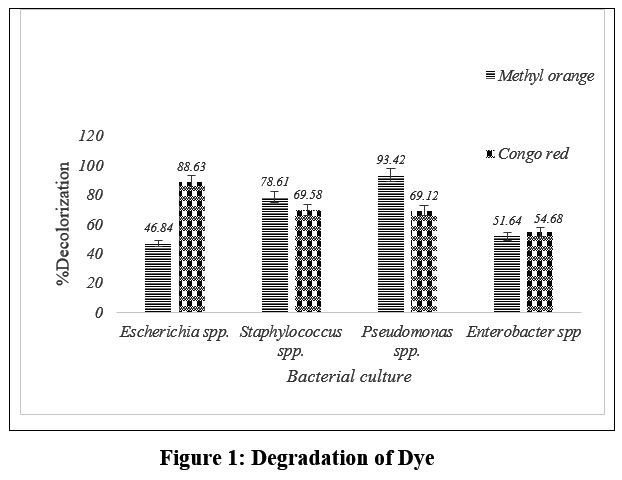 | Figure 1: Degradation of Dye
|
Optimization studies for methyl orange and Congo red
Effect of Temperature on Microbial Degradation of Dyes
The bacterial potential to degrade dyes may be affected by temperature. Temperature-dependent degradation of methyl orange and Congo red is shown in Figures 2 and 3. Each bacterial culture exhibited a different optimum degradation percentage at different temperatures. The highest degradation of methyl orange (94.36%) was observed with Escherichia spp. at room temperature & For, Congo red degradation it is 82.79% against Escherichia spp. at 30 °C.
The optimum degradation of methyl orange by Escherichia spp., Staphylococcus spp., Pseudomonas spp. and Enterobacter spp. was recorded at room temperature and 30 °C, with degradation percentages of 94.36%, 80.54%, 93.22% and 50.63%, respectively. Similarly, for Congo red, the optimum degradation by Escherichia spp., Staphylococcus spp., Pseudomonas spp. and Enterobacter spp. was recorded at 30 °C and room temperature, with degradation percentages of 82.79%, 77.65%, 63.52% and 55.39%, respectively. Among all strains, Enterobacter spp. showed the lowest degradation.
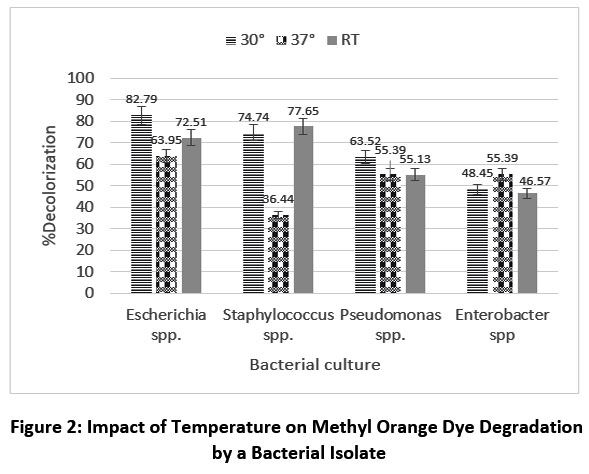 | Figure 2: Impact of Temperature on Methyl Orange Dye Degradation by a Bacterial Isolate
|
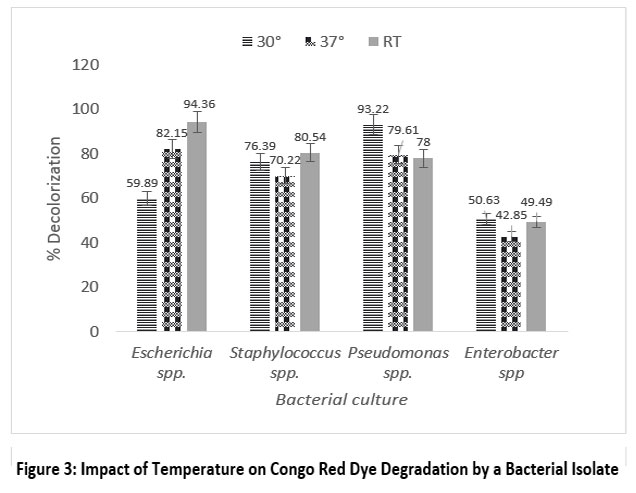 | Figure 3: Impact of Temperature on Congo Red Dye Degradation by a Bacterial Isolate
|
Effect of pH on Microbial Degradation of Dyes
The pH is an important factor affecting enzyme activity and the degradation potential of bacteria. The impact of pH on dye breakdown is shown. The optimum break down of methyl orange by Pseudomonas spp. occurred at pH 7, reaching 93.22%, while the optimum degradation of Congo red is by Staphylococcus spp. occurred at pH 9, reaching 89.94%.
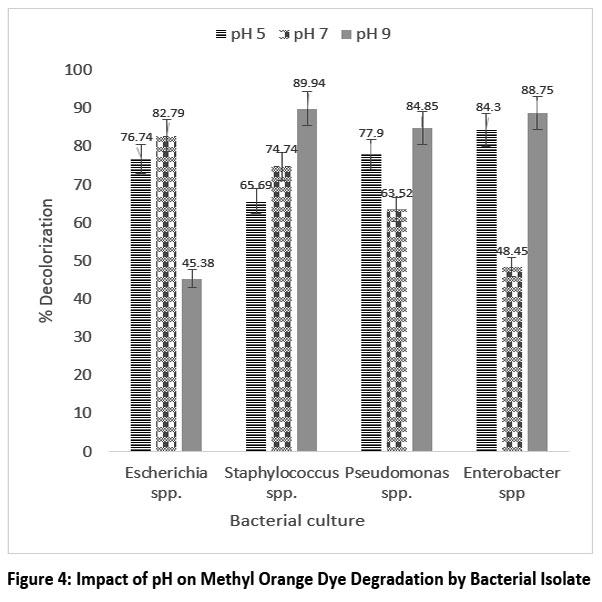 | Figure 4: Impact of pH on Methyl Orange Dye Degradation by Bacterial Isolate
|
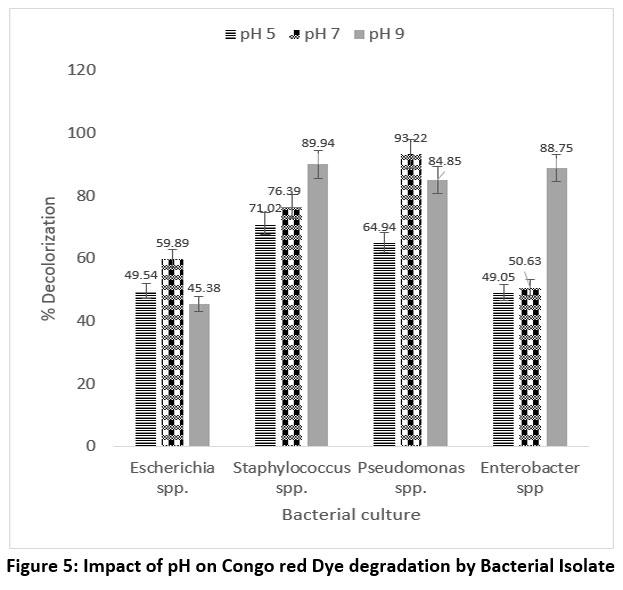 | Figure 5: Impact of pH on Congo red Dye degradation by Bacterial Isolate
|
Effect of Bacterial Consortium on Decolorization of Single and Mixed Dyes
The consortium degraded methyl orange faster than Congo red dye. After 24 hours, methyl orange degradation was 37.82%, while Congo red degradation was 15.75%. After 5 days, the degradation reached 95.70% and 94.43%, respectively. However, the consortium degraded the mixed dyes to 82.83% after incubation.
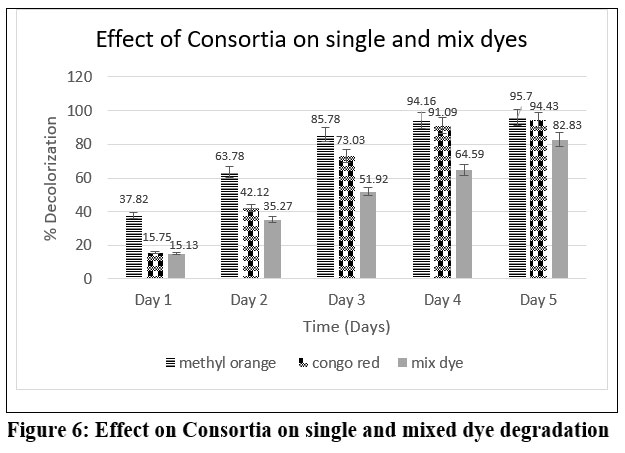 | Figure 6: Effect on Consortia on single and mixed dye degradation
|
FTIR analysis
The FTIR spectra comparison clearly shows dye has been degraded after treatment, as evidenced by the significant reduction in peak intensities in Sample 2. This confirms the effectiveness of the treatment process in breaking down the dye's molecular structure.
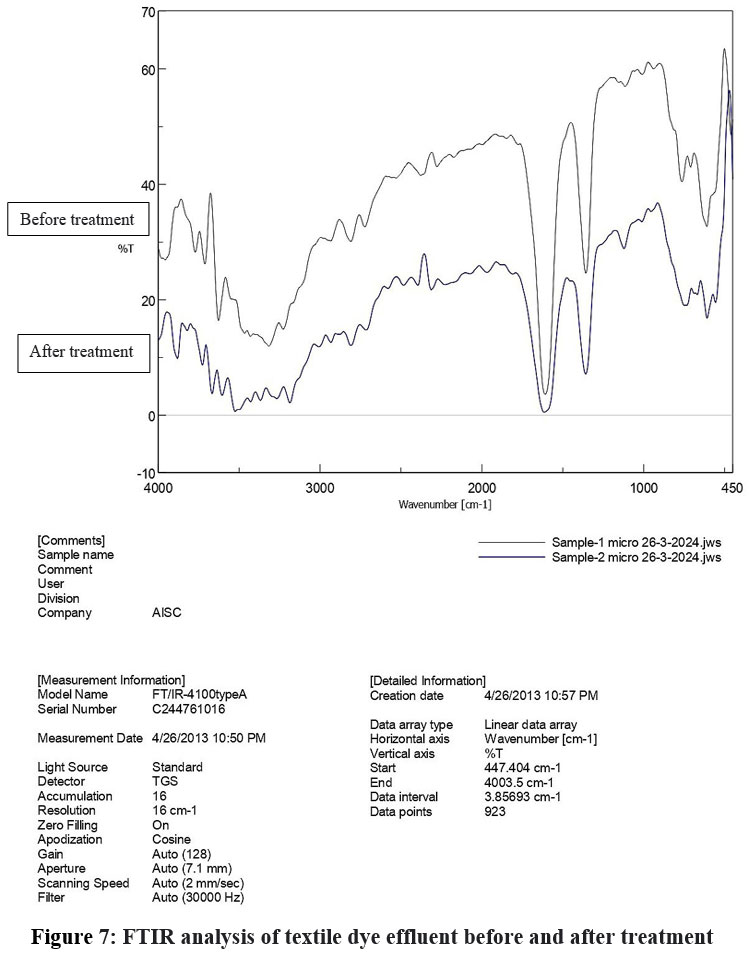 | Figure 7: FTIR analysis of textile dye effluent before and after treatment
|
Discussion
Escherichia spp. showed the highest degradation of methyl orange (94.36%) at room temperature and Congo red (82.79%) at 30 °C. Optimum methyl orange degradation for Staphylococcus spp., Pseudomonas spp. and Enterobacter spp. was 80.54%, 93.22% and 50.63%, respectively. For Congo red, optimum degradation by Staphylococcus spp., Pseudomonas spp. and Enterobacter spp. was 77.65%, 63.52% and 55.39%, with Enterobacter spp. showing the least degradation overall. Ikram, Muhammad, et al.,12 found that the optimum condition for Escherichia coli degradation of dye is 40 °C, in agreement with our observations. Kishor, Roop, et al.,13 reported that Pseudomonas aeruginosa shows 99% methyl orange degradation
At pH 7, Pseudomonas spp. achieved an optimum methyl orange degradation of 93.22%., while the highest degradation of Congo red (89.94%) was achieved by Staphylococcus spp. at pH 9. According to Shi, Yaqi, et al.,14 optimum degradation occurs within a pH range of 6–10. Similarly, Singh, R. P., et al.,15 reported maximum degradation at pH 7. Generally, dye decolorization is most effective under alkaline conditions.
The consortium degraded methyl orange faster than Congo red. After 24 hours, methyl orange degradation was 37.82%, whereas Congo red degradation was 15.75%. After 5 days, degradation reached 95.70% for methyl orange and 94.43% for Congo red. For mixed dyes, the consortium achieved 82.83% degradation after 5 days. Afrin, Sadia, et al.,16 reported similar results, showing that consortia degrade single dyes more effectively than mixtures. Decolorization efficiency depends on microbial adaptability, enzyme activity and microbial survival.
Conclusion
Rapid industrialization has led to the generation of various wastewaters contaminated with a range of toxic substances, including dyes, which contribute to environmental degradation and ecosystem imbalance. This situation has driven the search for new methods to treat textile dyes. The results indicate that bacteria have significant potential for use in the remediation of pollutants (dyes) from textile effluents. Among the four bacterial culture tested, Escherichia spp. and Pseudomonas spp. exhibited the highest decolorization efficiency at room temperature and pH 9 under static aerobic conditions.
Acknowledgement
The authors would like to express their sincere gratitude to Dr. Snehal Kulkarni, Head of the Department of Microbiology, Abeda Inamdar Senior College for her constant encouragement throughout the course of this research work. The authors are also deeply thankful to Dr. Shaila Bootwala, Principal, Abeda Inamdar Senior College for providing the necessary facilities, institutional support, and an encouraging academic environment that made this study possible.
Funding Sources:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflict of Interest
The author(s) do not have any conflict of interest.
Data Availability Statement
This statement does not apply to this article.
Ethics Statement
This research did not involve human participants, animal subjects, or any material that requires ethical approval.
Informed Consent Statement
This study did not involve human participants, and therefore, informed consent was not required.
Permission to reproduce material from other sources
Not Applicable
Author Contributions
Shilpa Pranav Tambe*: Conceptualization, Methodology, Supervision, Project Administration, Writing – Review & Editing.
Aarfiya Shaikh: Data Collection, Analysis, Writing – Original Draft.
Pranav Tambe: Visualization, Supervision, Project Administration, Writing – Review & Editing.
Shaikh Umair Ahmed: Data Collection, AnalysisJaspal Kaur Oberoi: Visualization, Supervision, Project Administration, Writing – Review & Editing.
References
- Al-Tohamy R, Ali SS, Li F, Okasha KM, Mahmoud YAG, Elsamahy T, Sun J. A critical review on the treatment of dye-containing wastewater: Ecotoxicological and health concerns of textile dyes and possible remediation approaches for environmental safety. Ecotoxicol Environ Saf. 2022;231:113160.
CrossRef - Jamee R, Siddique R. Biodegradation of synthetic dyes of textile effluent by microorganisms: an environmentally and economically sustainable approach. Eur J Microbiol Immunol. 2019;9(4):114-118.
CrossRef - Lellis B, Fávaro-Polonio CZ, Pamphile JA, Polonio JC. Effects of textile dyes on health and the environment and bioremediation potential of living organisms. Biotechnol Res Innov. 2019;3(2):275-290.
CrossRef - Saratale RG, Saratale GD, Chang JS, Govindwar SP. Bacterial decolorization and degradation of azo dyes: a review. J Taiwan Inst Chem Eng. 2011;42(1):138-157.
CrossRef - Senan RC, Abraham TE. Bioremediation of textile azo dyes by aerobic bacterial consortium. Biodegradation. 2004;15:275-280.
CrossRef - Siddique R, Ahmed HH. Isolation, identification and characterization of azo dye Reactive Violet 5R degrading bacterial strains from the textile sludge. J Bangladesh Acad Sci. 2018;41(2):136-143.
CrossRef - Shah MP, Patel KA, Nair SS, Darji AM. Isolation, identification and screening of dye decolorizing bacteria. Am J Microbiol Res. 2013;1(4):62-70.
CrossRef - Chequer FMD, Angeli JPF, Ferraz ERA, Tsuboy MS, Marcarini JC, Mantovani MS, de Oliveira DP. The azo dyes Disperse Red 1 and Disperse Orange 1 increase the micronuclei frequencies in human lymphocytes and in HepG2 cells. Mutat Res Genet Toxicol Environ Mutagen. 2009;676(1-2):83-86.
CrossRef - Ulson S.M.D.A.G., Bonilla K.A.S., de Souza A.A.U. Removal of COD and color from hydrolyzed textile azo dye by combined ozonation and biological treatment. J Hazard Mater. 2010;179(1-3):35-42.
CrossRef - Kamal IM, Abdeltawab NF, Ragab YM, Farag MA, Ramadan MA. Biodegradation, decolorization and detoxification of di-azo dye Direct Red 81 by halotolerant, alkali-thermo-tolerant bacterial mixed cultures. Microorganisms. 2022; 10(5):994.
CrossRef - Haque MM, Haque MA, Mosharaf MK, Marcus PK. Decolorization, degradation and detoxification of carcinogenic sulfonated azo dye methyl orange by newly developed biofilm consortia. Saudi J Biol Sci. 2021;28(1):793-804.
CrossRef - Ikram M, Naeem M, Zahoor M, Hanafiah MM, Oyekanmi AA, Ullah R, Gulfam N. Biological degradation of the azo dye basic orange 2 by Escherichia spp.: A sustainable and ecofriendly approach for the treatment of textile wastewater. Water. 2022;14(13):2063.
CrossRef - Kishor R, Purchase D, Saratale GD, Ferreira LFR, Hussain CM, Mulla SI, Bharagava RN. Degradation mechanism and toxicity reduction of methyl orange dye by a newly isolated bacterium Pseudomonas spp. MZ520730. J Water Process Eng. 2021;43:102300.
CrossRef - Shi Y, Yang Z, Xing L, Zhang X, Li X, Zhang D. Recent advances in the biodegradation of azo dyes. World J Microbiol Biotechnol. 2021;37:1-18.
CrossRef - Singh RP, Singh PK, Singh RL. Bacterial decolorization of textile azo dye acid orange by Staphylococcus hominis RMLRT03. Toxicol Int. 2014;21(2):160.
CrossRef - Afrin S, Shuvo HR, Sultana B, Islam F, Rus'd AA, Begum S, Hossain MN. The degradation of textile industry dyes using the effective bacterial consortium. Heliyon. 2021;7(10):e08102.
CrossRef






